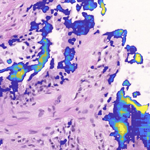
RationAI: showcase visualization
You can find interactive visualizations of our current research here.
Automated annotations of epithelial cells and stroma in hematoxylin–eosin-stained whole-slide images using cytokeratin re-staining
Abstract
The diagnosis of solid tumors of epithelial origin (carcinomas) represents a major part of the workload in clinical histopathology.
Carcinomas consist of malignant epithelial cells arranged in more or less cohesive clusters of variable size and shape, together with stromal cells,
extracellular matrix, and blood vessels. Distinguishing stroma from epithelium is a critical component of artificial intelligence (AI) methods developed to detect and analyze carcinomas.
In this paper, we propose a novel automated workflow that enables large-scale guidance of AI methods to identify the epithelial component. The workflow is based on re-staining existing
hematoxylin and eosin (H&E) formalin-fixed paraffin-embedded sections by immunohistochemistry for cytokeratins, cytoskeletal components specific to epithelial cells. Compared to existing methods,
clinically available H&E sections are reused and no additional material, such as consecutive slides, is needed. We developed a simple and reliable method for automatic alignment to generate masks
denoting cytokeratin-rich regions, using cell nuclei positions that are visible in both the original and the re-stained slide. The registration method has been compared to state-of-the-art methods
for alignment of consecutive slides and shows that, despite being simpler, it provides similar accuracy and is more robust. We also demonstrate how the automatically generated masks can be used
to train modern AI image segmentation based on U-Net, resulting in reliable detection of epithelial regions in previously unseen H&E slides. Through training on real-world material available
in clinical laboratories, this approach therefore has widespread applications toward achieving AI-assisted tumor assessment directly from scanned H&E sections. In addition, the re-staining method
will facilitate additional automated quantitative studies of tumor cell and stromal cell phenotypes.
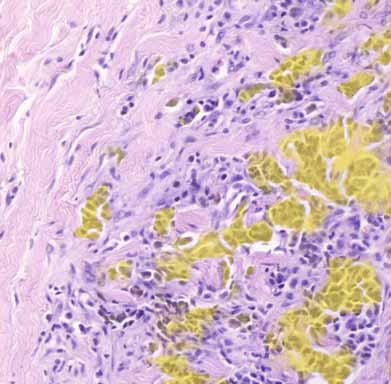
Illustration of WSIs available online including U-Net computed epithelial mask in colon and breast carcinoma.
epithelial mask
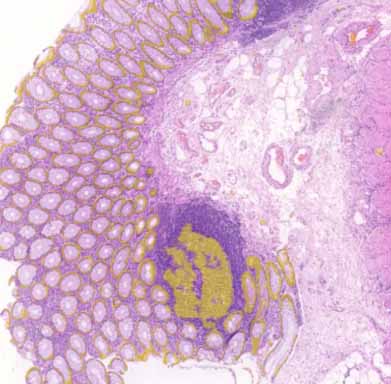
epithelial mask
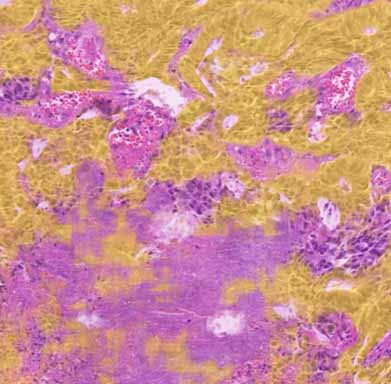
epithelial mask
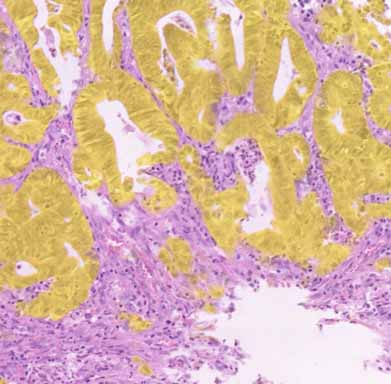
epithelial mask
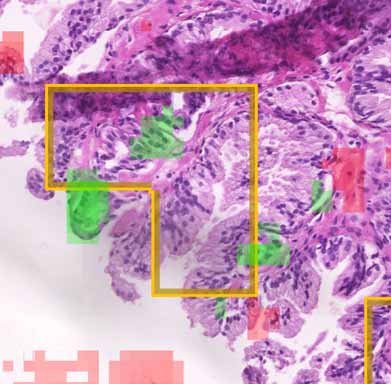
Automated cancer detection in a tissue scan
This is an example from the IEEE video contest: a tissue scan
where cancerous regions are being detected using a neural network. There are three available data layers you can view:
- Annotation: original annotation made by pathologist
- Probability: rectangular areas where the network encodes the probability of a cancer in the rectangle opacity
- Explainability: by covering and evaluating aforementioned rectangle parts we detect what areas are contributing to the
decision and thus showing where the network thinks it can see positive (green) or negative (red) cases.
Interactive detection example.
VGG16
U-Net (with a mask)

XOpat: a WSI Viewer
Introduction
The whole slide imaging (WSI), also known as virtual microscopy, refers to a cost-effective method to digitize a whole glass slides, for example with a stained tissue sample. The technology emerged in the late 1990s' and it is considered as a disruptive technology that enabled new ways for diagnostic, educational and research purposes in digital pathology.
Digitized whole slide images (thereof referred to as images) are typically multi-resolution data accessible through specialized image viewers.
Illustration of fast neural network results visualization. There are many parameters to play with and simple, effective visualization on the whole-slide scale helps to spot invalid configurations or mistakes quickly. It also introduces uniform way of communication and screen alignment with many other features (plugins) available.
wrong slicing on the input data
a library is in BGR mode by default